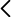The genomic analysis has been carried out by the Joint Unit on Infection and Public Health of the University of Valencia and the Foundation for the Promotion of Health and Biomedical Research of the Valencian Community Fisabio, the Research Group in Molecular Epidemiology, both led by Fernando González Candelas, Professor of Genetics and researcher at the Institute for Integrative Systems Biology (I2SysBio) of the University of Valencia and CSIC, and the Fisabio Sequencing and Bioinformatics Service coordinated by Giuseppe d’Auria and Llúcia Martínez.
The Valencian research teams have managed to sequence the complete genome of three samples from patients infected with COVID-19 from the Microbiology laboratory of the University Clinical Hospital of Valencia, the first sequences of the SARS-CoV2 virus obtained in Spain. This complete genome sequencing adds to the effort being made worldwide in all laboratories to find out what the transmission routes have been and how the different lineages of the virus have spread.
“The primary objective is to identify more reliably the foci and transmission chains of the coronavirus. The main conclusion of the analysis of the first samples is that the strains come from different transmission routes”, explains the researcher Fernando González Candelas.
The sequences are already accessible in the GISAID Initiative database, a public consortium dedicated to the study of the influenza virus (https://www.gisaid.org); the Nextstrain platform, which allows visualising the spatial and temporal progression of the pandemic from the more than 500 genomes deposited since last December by 40 countries (https://nextstrain.org); and the Genbank database of genetic sequences from the NIH (National Institutes of Health of the United States) (https://www.ncbi.nlm.nih.gov/genbank).
The groups made up of research staff specialised in virology, epidemiology and bioinformatics have determined that one of the strains analysed is related to other European strains (from Italy, Germany, Luxembourg, France, Scotland, the Netherlands, etc.). “The next step will be to analyse sequences of more samples of patients from the hospitals of the Valencian Community in order to check the links between them and with the transmission chains established by the epidemiology specialist personnel”, explains Fernando González.
The analysis of viral genomes allows us to know the ways in which the virus has entered our community and how it is transmitted at this time, which will help health authorities to better control the spread of the virus in our community. In addition, the sequencing of the virus genome allows us to know the mutations that the virus has suffered since the epidemic began and the conclusion that has been reached after the analysis carried out is that, until now, no mutation associated with increased virulence, lethality, or some interesting property from a clinical point of view.
The samples have been sequenced using MinIOn, a third-generation sequencer from Oxford Nanopore Technologies. The infrastructure used in the research has been possible thanks to the co-financing of the European Union through the Operational Program of the European Regional Development Fund (ERDF) of the Valencian Community 2014-2020.
More information:
http://fisabio.san.gva.es/ca/inicio
Photo captions:
1. Sequencing Team Group
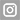
2. Image of a fragment of the SARS-CoV-2 virus sequences analysed. Blue shows a mutation of the virus with respect to the reference genome.
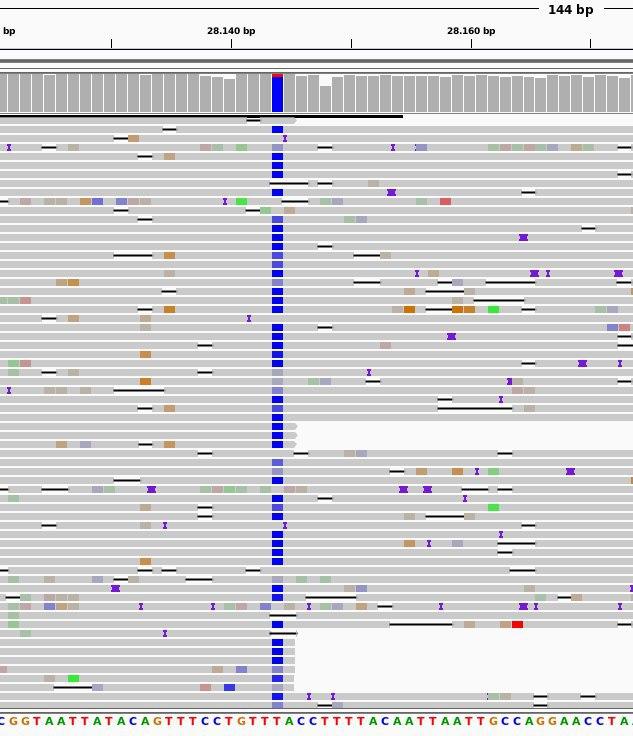
3. Phylogeny of the SARS-CoV-2 coronavirus. The arrows indicate the lineages of the viruses isolated in Valencia.
Images: