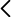BactGem
Title: Genomic-scale metabolic models and bacterial reverse ecology: application to bacteria of economic interest
Research groups: Biotechnology and Synthetic Biology and Systems Metabolic Engineering
This project will contribute to the identification of the molecular mechanism of action of probiotic microorganisms, based on the identification of the synthesis pathways of molecular patterns relevant to their functional action. A double approach is proposed: computational and experimental. Thus, a series of metabolic models will be developed at a genomic scale (GEM) from the annotated sequences of bifidobacteria genomes. This will allow us to obtain the functional metabolic reconstructions and to analyze, among other aspects, growth capacities, carbon source utilization or essential factor requirements. Likewise, the production of metabolites of interest can be simulated and analyzed. This knowledge will serve as a basis for the characterization of the mechanisms of probiotic action. In this way, it will be possible to obtain loss of function mutants in the microorganisms of interest, through the selective deletion of candidate genes based on the comparative analysis of their genomes and associated metabolic models. Finally, by means of reverse ecology analysis, physiological intervention strategies will be evaluated. The general strategy of reverse ecology is also applicable to microorganisms that present very slow growth kinetics, since it allows us to optimize, first in silico and then in vitro, the culture conditions. As a proof of concept, we propose the design of a minimum culture medium for the growth of the phytopathogen Xylella fastidosa.
Ref.PIE202020E120
Juli Peretó
Patricia Álvarez, Alba Arévalo, Paola Corbín, José Luis García López, Daniel Ramón, Marta Tortajada.
Consell Superior d'Investigacions Científiques (CSIC).