Projects
Folding, insertion and dimerization of transmembrane protein fragments
PI: Víctor Lórenz-Fonfría
Our aim is to understand, at the molecular level, the mechanism and the dynamics of folding, insertion and dimerization of transmembrane (TM) helices in lipidic membranes. To reach that goal, we are adding optical control to the processes we want to understand.
We are primarily using photopeptides and photolipids (i.e., peptides and lipids chemically engineered to respond to light), and photoacids (molecules with a pKa that responds to light). As model systems to study folding, insertion and dimerization of TM helices we are using monomeric helical peptides with a pH-sensitive membrane insertion, and TM fragments of proteins that dimerize by helix-helix interaction. We chacterize how our systems respond to light by means of state-of-the-art IR spectroscopic methods, complemented by other spectroscopies (UV-vis, CD, fluorescence and Raman) and molecular dynamics simulations.
We are primarily using photopeptides and photolipids (i.e., peptides and lipids chemically engineered to respond to light), and photoacids (molecules with a pKa that responds to light). As model systems to study folding, insertion and dimerization of TM helices we are using monomeric helical peptides with a pH-sensitive membrane insertion, and TM fragments of proteins that dimerize by helix-helix interaction. We chacterize how our systems respond to light by means of state-of-the-art IR spectroscopic methods, complemented by other spectroscopies (UV-vis, CD, fluorescence and Raman) and molecular dynamics simulations.
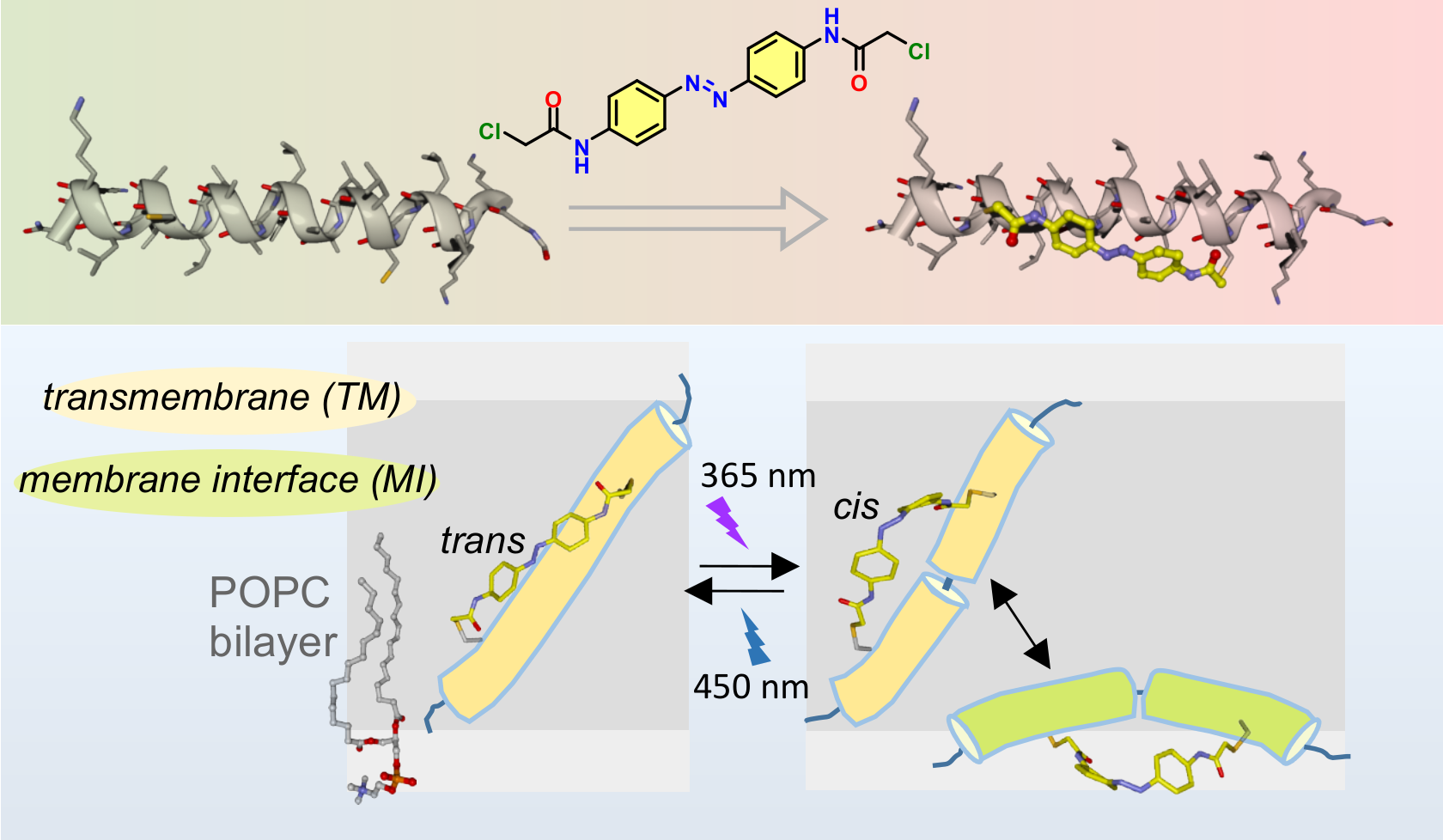
A chemically engineered peptide which inserts and exits the hydrophobic region of a lipidic membrane in response to light
- Diego Sampredro , Department of Chemistry, Universidad de La Rioja
- Joachim Heberle , Department of Physics | AG Heberle, Freie Universität Berlin
- PID2019-106103GB-I00, MINECO / MICINN
- BFU2016-76805-P, MINECO / MICINN
Vibrational pH-sensitive probes
PI: Víctor Lórenz-Fonfría
We are developing the use of biological buffers as pH-sensitive vibrational probes, aiming to solve some of the limiations of traditional pH-probes based on chromophores and fluorophores. The pH-vibrational probes will be used to track with high fidelity protonation changes during the functional mechanism of membrane proteins by time-resolved vibrational spectroscopies, as well as to obtain pH maps of gigant unilamellar vesicles with vibrational microscopies.
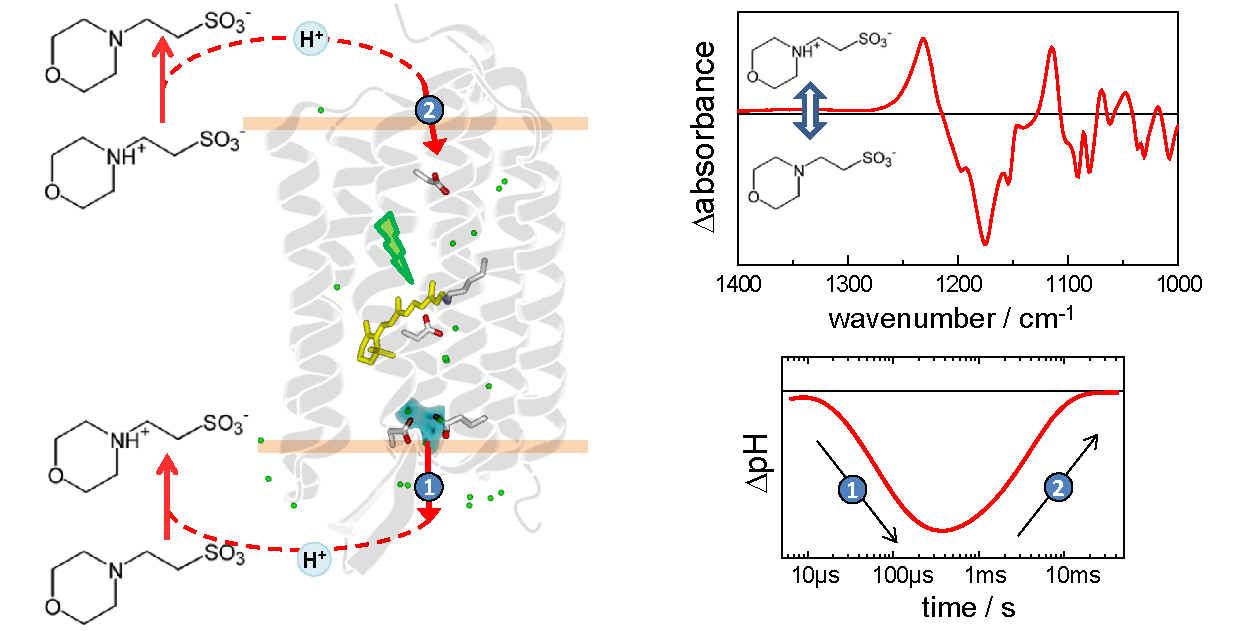
- Joachim Heberle , Department of Physics | AG Heberle, Freie Universität Berlin
- BFU2017‐91559‐EXP, MICINN


