Collaborations
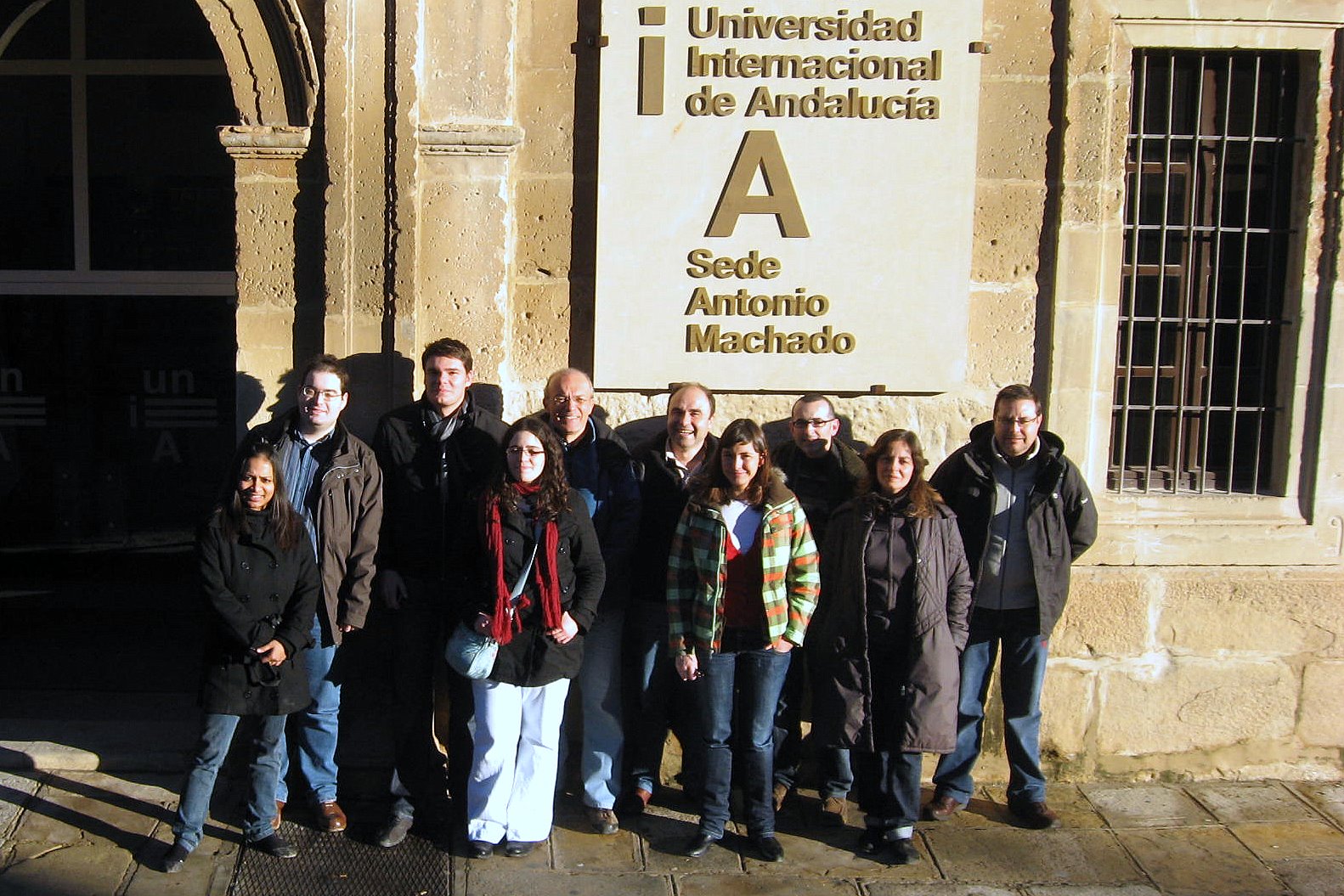

We collaborate with researchers below:
- Dr. Joaquin Moreno, Dept. of Biochemistry and Molecular Biology, University of Valencia. Collaboration initiated in 2006. We work in theoretical studies of regulation of gene expression and genomic data analysis. Dr. Moreno is a specialist in the development of mathematical algorithms to calculate, from genomic data kinetic expression, unsteady state variables.
- Dr. Per Sunnerhagen and Dr. Jonas Warringer. Dpt. of Cell and Molecular Biology. Lundberg Laboratory, Göteborg University, Sweden. Collaboration was initiated in 2008 to study genomic translation rate and study the function of various RNA-binding proteins in regulating translation rates in response to osmotic stress. This collaboration has intensified after a stay of six months as a visiting researcher of Dr. Alepuz in the laboratory of Drs Sunnerhagen and Warringer in 2011. We are currently awaiting the resolution of a joint request to the Swedish Foundation for International Cooperation in Research and High Education (STINT). Additionally, Dr. Warringer has granted a short stay of three months in our laboratory for 2012-13 funded by the program "attract Talent" of the University of Valencia. In the laboratories of Drs Sunnerhagen Warringer in Sweden and has systems for Bioscreen quantitative phenotypic analysis, and robotic systems for fluorescent signal readings in 96-well plates. These systems are used for screening mutant quantitative scale.
- Dr. Alejandro Ferrando. IBM-PC (CSIC), Valencia. Collaboration initiated in 2011 to study the translational control of mRNAs involved in flowering by Tif51 translation factor (eIF5A). Dr. Ferrando has performed in our laboratory analysis of mutant plants in translation through the fractionator Tif51 polysomes.
- Dr. Gustav Ammerer and Dr. Aurora Zuzuarregui, Dept. of Biochemistry and Molecular Cell Biology, Max F. Perutz Laboratories, University of Vienna, Vienna, Austria. The group of Dr. Ammerer has perfected the method M-Track to capture transient protein interactions. Together, we are currently analyzing interactions between components of the HOG signaling pathway at the level of signal reception at the plasma membrane, and interactions between components of the path and the translation machinery. M-Track to have the antibodies specifically designed for this method by the Vienna group.
-Dr. Olga Calvo (USAL) in the study of genome-wide functions SUB1 gene of S. cerevisiae by studying the sub1 mutant transcritpome with "tiling arrays."
-Dr. Sebastian Chavez (U.S.). With the group of Prof. Chavez we have been collaborating since 2005. We had 4 projects all coordinated in the field of the study of transcriptional elongation using molecular genetic techniques and genomics. We have co-published six original and review articles and we have performed multiple stays of doctoral or senior researchers of one of the laboratories in the other group.
-Dr. Francisco Navarro (University of Jaén). As with the group of Dr. Chavez have had four coordinated projects. This group is expert in genetics and molecular structure of RNA polymerase II. So far it has published an article in the journal Genetics..
- Dr. Enrique Herrero (UDL), Dr. Jesus Pla (UCM) and Dr. Joachim Ernst (Heinrich-Heine-Universität Düsseldorf) in the genome of transcriptional and post-transcriptional responses to oxidative stress in C. albicans, within a project PATHOGENOMICS (ERA-Net).
-Dr. Joaquin Ariño (UAB) in the genome of transcriptional and post-transcriptional responses to alkaline pH stress in S. cerevisiae.
- Dr. Jesus de la Cruz (U.S.) in the genome-wide study of the phenotypes caused by a mutation in the gene RPL3 of S. cerevisiae.
-Dr. Mordechai Choder (Technion, Israel) in the genome-wide study of the respective influences of the transcription machinery and degradation of mRNAs. This collaboration has confirmed the hypothesis: the existence of homeostasis in the amount of mRNA in the cell. Our results suggest that there are global compensation mechanisms and cross-talk between the machinery of synthesis and degradation of mRNAs.