-
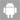
eIF5A controls mitoprotein import by relieving ribosome stalling at TIM50 translocase mRNA
Barba-Aliaga M*, Bernal V, Rong C, Volfbeyn ME, Zhang K, Zid BM*, Alepuz P*. J Cell Biol. 2024 Dec 2;223(12):e202404094.
(2024). ArticleJournal of Cell Biology (JCB). No.v. 223, Issue 12
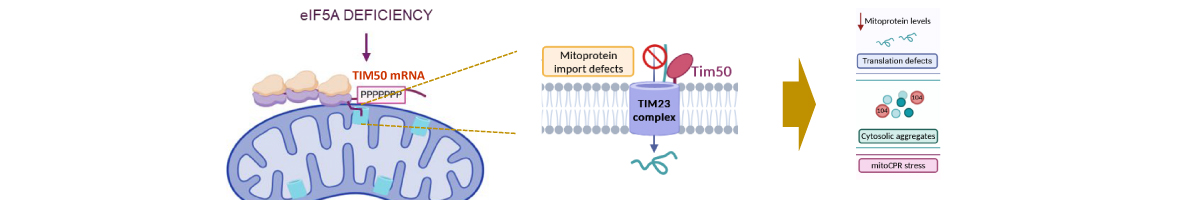
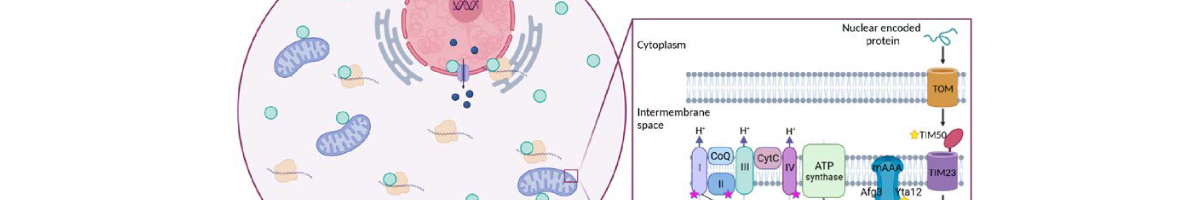
Efficient import of nuclear-encoded proteins into mitochondria is crucial for proper mitochondrial function. The conserved translation factor eIF5A binds ribosomes, alleviating stalling at polyproline-encoding sequences. eIF5A impacts mitochondrial function across species, though the precise molecular mechanism is unclear. We found that eIF5A depletion in yeast reduces the translation and levels of the TCA cycle and oxidative phosphorylation proteins. Loss of eIF5A causes mitoprotein precursors to accumulate in the cytosol and triggers a mitochondrial import stress response. We identify an essential polyproline protein as a direct target of eIF5A: the mitochondrial inner membrane protein...
Efficient import of nuclear-encoded proteins into mitochondria is crucial for proper mitochondrial function. The conserved translation factor eIF5A binds ribosomes, alleviating stalling at polyproline-encoding sequences. eIF5A impacts mitochondrial function across species, though the precise molecular mechanism is unclear. We found that eIF5A depletion in yeast reduces the translation and levels of the TCA cycle and oxidative phosphorylation proteins. Loss of eIF5A causes mitoprotein precursors to accumulate in the cytosol and triggers a mitochondrial import stress response. We identify an essential polyproline protein as a direct target of eIF5A: the mitochondrial inner membrane protein and translocase component Tim50. Thus, eIF5A controls mitochondrial protein import by alleviating ribosome stalling along Tim50 mRNA at the mitochondrial surface. Removal of polyprolines from Tim50 partially rescues the mitochondrial import stress response and translation of oxidative phosphorylation genes. Overall, our findings elucidate how eIF5A impacts the mitochondrial function by promoting efficient translation and reducing ribosome stalling of co-translationally imported proteins, thereby positively impacting the mitochondrial import process.
Read more HidePMID: 39509053
*corresponding author
DOI: 10.1083/jcb.202404094 -

eIF5A promotes +1 programmed ribosomal frameshifting in Euplotes octocarinatus
Xiao Y, Li J, Wang R, Fan Y, Han X, Fu Y, Alepuz P, Wang W, Liang A*. Int J Biol Macromol. 2024 Jan;254(Pt 1):127743
(2024). ArticleInternational Journal of Biological Macromolecules. No.Volume 254, Part 1, January 2024, 127743
Programmed ribosomal frameshifting (PRF) exists in all branches of life that regulate gene expression at the translational level. The single-celled eukaryote Euplotes exhibit high frequency of PRF. However, the molecular mechanism of modulating Euplotes PRF remains largely unknown. Here, we identified two novel eIF5A genes, eIF5A1 and eIF5A2, in Euplotes octocarinatus and found that the Eo-eIF5A2 gene requires a −1 PRF to produce complete protein product. Although both Eo-eIF5As showed significant structural similarity with yeast eIF5A, neither of them could functionally replace yeast eIF5A. Eo-eIF5A knockdown inhibited +1 PRF of the η-tubulin gene. Using an in vitro reconstituted...
Programmed ribosomal frameshifting (PRF) exists in all branches of life that regulate gene expression at the translational level. The single-celled eukaryote Euplotes exhibit high frequency of PRF. However, the molecular mechanism of modulating Euplotes PRF remains largely unknown. Here, we identified two novel eIF5A genes, eIF5A1 and eIF5A2, in Euplotes octocarinatus and found that the Eo-eIF5A2 gene requires a −1 PRF to produce complete protein product. Although both Eo-eIF5As showed significant structural similarity with yeast eIF5A, neither of them could functionally replace yeast eIF5A. Eo-eIF5A knockdown inhibited +1 PRF of the η-tubulin gene. Using an in vitro reconstituted translation system, we found that hypusinated Eo-eIF5A (Eo-eIF5AH) can promote +1 PRF at the canonical AAA_UAA frameshifting site of Euplotes. The results showed eIF5A is a novel trans-regulator of PRF in Euplotes and has an evolutionary conserved role in regulating +1 PRF in eukaryotes.
Read more HidePMID: 38287569
*corresponding author
DOI: 10.1016/j.ijbiomac.2023.127743 -
SUMOylation modulates eIF5A activities in both yeast and pancreatic ductal adenocarcinoma cells
Seoane R, Lama-Díaz T, Romero AM, El Motiam A, Martínez-Férriz A, Vidal S, Bouzaher YH, Blanquer M, Tolosa RM, Castillo Mewa J, Rodríguez MS, García-Sastre A, Xirodimas D, Sutherland JD, Barrio R, Alepuz P, Blanco MG, Farràs R, Rivas C*.
(2024). ArticleCellular & Molecular Biology Letters. No.29, Article number: 15 (2024)
The eukaryotic translation initiation protein eIF5A is a highly conserved and essential factor that plays a critical role in different physiological and pathological processes including stress response and cancer. Different proteomic studies suggest that eIF5A may be a small ubiquitin-like modifier (SUMO) substrate, but whether eIF5A is indeed SUMOylated and how relevant is this modification for eIF5A activities are still unknown.
PMID: 38229033
*corresponding author
DOI: 10.1186/s11658-024-00533-5 -
Repositorio Datos: ProteomeXchange Consortium
M. Barba-Aliaga, V. Bernal, C. Rong, B. Zid, P. Alepuz
(2023). Recurs electrònicJ Cell Biol. 2024. No.223(12):e202404094
Acceso dataset: PXD043905. Tipo de datos: Proteómica
-
The activator/repressor Hap1 binds to the yeast eIF5A-encoding gene TIF51A to adapt its expression to the mitochondrial functional status
Barba-Aliaga M*, Alepuz P*. FEBS Lett. 2022 Jul;596(14):1809-1826
(2022). ArticleFEBS letters. No.Jul;596(14):1809-1826
Mitochondrial activity adapts to cellular energetic and metabolic demands; its dysfunction is a hallmark of ageing and many human diseases. The evolutionarily conserved translation elongation factor eIF5A is involved in maintaining mitochondrial function. In humans, eIF5A is encoded by two highly homologous but differentially expressed genes; in yeast, these are TIF51A and TIF51B. We show that yeast transcription factor Hap1 constitutively binds to the TIF51A promoter to activate its expression under respiration, but represses its expression under nonrespiration conditions by recruiting the corepressor Tup1. Hap1 indirectly regulates TIF51B expression by binding to and activating the TIF51B...
Mitochondrial activity adapts to cellular energetic and metabolic demands; its dysfunction is a hallmark of ageing and many human diseases. The evolutionarily conserved translation elongation factor eIF5A is involved in maintaining mitochondrial function. In humans, eIF5A is encoded by two highly homologous but differentially expressed genes; in yeast, these are TIF51A and TIF51B. We show that yeast transcription factor Hap1 constitutively binds to the TIF51A promoter to activate its expression under respiration, but represses its expression under nonrespiration conditions by recruiting the corepressor Tup1. Hap1 indirectly regulates TIF51B expression by binding to and activating the TIF51B repressor genes ROX1 and MOT3 under respiration and repressing them under nonrespiration. Thus, the levels of eIF5A isoforms are adapted to the mitochondrial functional status.
Read more HidePMID: 35490374
*corresponding author
DOI: 10.1002/1873-3468.14366 -
Role of eIF5A in Mitochondrial Function
Barba-Aliaga M*, Alepuz P*.
(2022). ArticleInternational Journal of Molecular Sciences (Int. J. Mol. Sci.). No.2022, 23(3), 1284
The eukaryotic translation initiation factor 5A (eIF5A) is an evolutionarily conserved protein that binds ribosomes to facilitate the translation of peptide motifs with consecutive prolines or combinations of prolines with glycine and charged amino acids. It has also been linked to other molecular functions and cellular processes, such as nuclear mRNA export and mRNA decay, proliferation, differentiation, autophagy, and apoptosis. The growing interest in eIF5A relates to its association with the pathogenesis of several diseases, including cancer, viral infection, and diabetes. It has also been proposed as an anti-aging factor: its levels decay in aged cells, whereas increasing levels of...
The eukaryotic translation initiation factor 5A (eIF5A) is an evolutionarily conserved protein that binds ribosomes to facilitate the translation of peptide motifs with consecutive prolines or combinations of prolines with glycine and charged amino acids. It has also been linked to other molecular functions and cellular processes, such as nuclear mRNA export and mRNA decay, proliferation, differentiation, autophagy, and apoptosis. The growing interest in eIF5A relates to its association with the pathogenesis of several diseases, including cancer, viral infection, and diabetes. It has also been proposed as an anti-aging factor: its levels decay in aged cells, whereas increasing levels of active eIF5A result in the rejuvenation of the immune and vascular systems and improved brain cognition. Recent data have linked the role of eIF5A in some pathologies with its function in maintaining healthy mitochondria. The eukaryotic translation initiation factor 5A is upregulated under respiratory metabolism and its deficiency reduces oxygen consumption, ATP production, and the levels of several mitochondrial metabolic enzymes, as well as altering mitochondria dynamics. However, although all the accumulated data strongly link eIF5A to mitochondrial function, the precise molecular role and mechanisms involved are still unknown. In this review, we discuss the findings linking eIF5A and mitochondria, speculate about its role in regulating mitochondrial homeostasis, and highlight its potential as a target in diseases related to energy metabolism.
Read more HidePMID: 35163207 Review.
*corresponding author
(This article belongs to the Special Issue Mitochondria in Human Diseases)
DOI: 10.3390/ijms23031284 -
Repositorio Datos: Gene expression omnibus (GEO)
García-Martínez J, Pérez-Martínez ME, Pérez-Ortín JE, Alepuz P
(2021). Recurs electrònicRNA Biol. No.18(10):1458-1474
Acceso dataset: GSE151736. Tipo de datos: Genomic Run On (RNA expresión)
-
Xrn1 influence on gene transcription results from the combination of general effects on elongating RNA pol II and gene-specific chromatin configuration
Begley V, Jordán-Pla A, Peñate X, Garrido-Godino AI, Challal D, Cuevas-Bermúdez A, Mitjavila A, Barucco M, Gutiérrez G, Singh A, Alepuz P, Navarro F, Libri D, Pérez-Ortín JE, Chávez S.
(2021). ArticleRNA Biol. No.Volume 18 - Issue 9, pp. 1310-1323
eIF5A is an essential and evolutionary conserved translation elongation factor, which has recently been proposed to be required for the translation of proteins with consecutive prolines. The binding of eIF5A to ribosomes occurs upon its activation by hypusination, a modification that requires spermidine, an essential factor for mammalian fertility that also promotes yeast mating. We show that in response to pheromone, hypusinated eIF5A is required for shmoo formation, localization of polarisome components, induction of cell fusion proteins, and actin assembly in yeast. We also show that eIF5A is required for the translation of Bni1, a proline-rich formin involved in polarized growth during...
eIF5A is an essential and evolutionary conserved translation elongation factor, which has recently been proposed to be required for the translation of proteins with consecutive prolines. The binding of eIF5A to ribosomes occurs upon its activation by hypusination, a modification that requires spermidine, an essential factor for mammalian fertility that also promotes yeast mating. We show that in response to pheromone, hypusinated eIF5A is required for shmoo formation, localization of polarisome components, induction of cell fusion proteins, and actin assembly in yeast. We also show that eIF5A is required for the translation of Bni1, a proline-rich formin involved in polarized growth during shmoo formation. Our data indicate that translation of the polyproline motifs in Bni1 is eIF5A dependent and this translation dependency is lost upon deletion of the polyprolines. Moreover, an exogenous increase in Bni1 protein levels partially restores the defect in shmoo formation seen in eIF5A mutants. Overall, our results identify eIF5A as a novel and essential regulator of yeast mating through formin translation. Since eIF5A and polyproline formins are conserved across species, our results also suggest that eIF5A-dependent translation of formins could regulate polarized growth in such processes as fertility and cancer in higher eukaryotes.
Read more Hide DOI: 10.1080/15476286.2020.1845504 -
Hypusinated eIF5A is required for the translation of collagen
Barba-Aliaga M, Mena A, Espinoza V, Apostolova N, Costell M, Alepuz P*.
(2021). ArticleJournal of Cell Science. No.v. 134, Issue 18. Sep 2021
Translation of mRNAs that encode peptide sequences with consecutive prolines (polyproline) requires the conserved and essential elongation factor eIF5A to facilitate the formation of peptide bonds. It has been shown that, upon eIF5A depletion, yeast ribosomes stall in polyproline motifs, but also in tripeptide sequences that combine proline with glycine and charged amino acids. Mammalian collagens are enriched in putative eIF5A-dependent Pro-Gly-containing tripeptides. Here, we show that depletion of active eIF5A in mouse fibroblasts reduced collagen type I α1 chain (Col1a1) content, which concentrated around the nuclei. Moreover, it provoked the upregulation of endoplasmic reticulum (ER)...
Translation of mRNAs that encode peptide sequences with consecutive prolines (polyproline) requires the conserved and essential elongation factor eIF5A to facilitate the formation of peptide bonds. It has been shown that, upon eIF5A depletion, yeast ribosomes stall in polyproline motifs, but also in tripeptide sequences that combine proline with glycine and charged amino acids. Mammalian collagens are enriched in putative eIF5A-dependent Pro-Gly-containing tripeptides. Here, we show that depletion of active eIF5A in mouse fibroblasts reduced collagen type I α1 chain (Col1a1) content, which concentrated around the nuclei. Moreover, it provoked the upregulation of endoplasmic reticulum (ER) stress markers, suggesting retention of partially synthesized collagen 1 (Col1) in the ER. We confirmed that eIF5A is needed for heterologous collagen synthesis in yeast and, using a double luciferase reporter system, showed that eIF5A depletion interrupts translation at Pro-Gly collagenic motifs. A dramatically lower level of Col1a1 protein was also observed in functional eIF5A-depleted human hepatic stellate cells treated with the profibrotic cytokine TGF-β1. In sum, our results show that collagen expression requires eIF5A and imply its potential as a target for regulating collagen production in fibrotic diseases.
Read more HidePMID: 34447991
*corresponding author
DOI: 10.1242/jcs.258643 -
Eukaryotic RNA Polymerases: The Many Ways to Transcribe a Gene
Barba-Aliaga M, Alepuz P, Pérez-Ortín JE*.
(2021). ArticleFrontiers in Molecular Biosciences. No.v. 8 - 2021
In eukaryotic cells, three nuclear RNA polymerases (RNA pols) carry out the transcription from DNA to RNA, and they all seem to have evolved from a single enzyme present in the common ancestor with archaea. The multiplicity of eukaryotic RNA pols allows each one to remain specialized in the synthesis of a subset of transcripts, which are different in the function, length, cell abundance, diversity, and promoter organization of the corresponding genes. We hypothesize that this specialization of RNA pols has conditioned the evolution of the regulatory mechanisms used to transcribe each gene subset to cope with environmental changes. We herein present the example of the homeostatic regulation...
In eukaryotic cells, three nuclear RNA polymerases (RNA pols) carry out the transcription from DNA to RNA, and they all seem to have evolved from a single enzyme present in the common ancestor with archaea. The multiplicity of eukaryotic RNA pols allows each one to remain specialized in the synthesis of a subset of transcripts, which are different in the function, length, cell abundance, diversity, and promoter organization of the corresponding genes. We hypothesize that this specialization of RNA pols has conditioned the evolution of the regulatory mechanisms used to transcribe each gene subset to cope with environmental changes. We herein present the example of the homeostatic regulation of transcript levels versus changes in cell volume. We propose that the diversity and instability of messenger RNAs, transcribed by RNA polymerase II, have conditioned the appearance of regulatory mechanisms based on different gene promoter strength and mRNA stability. However, for the regulation of ribosomal RNA levels, which are very stable and transcribed mainly by RNA polymerase I from only one promoter, different mechanisms act based on gene copy variation, and a much simpler regulation of the synthesis rate.
Read more HidePMID: 33968992 Review.
*corresponding author
DOI: 10.3389/fmolb.2021.663209