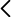Title: Genomics and the evolution of bacteria resistant to antibiotics: from molecular epidemiology to phylogenomics
Research Group: Molecular Epidemiology
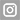
Antibiotic resistance represents one of the greatest threats to global public health. Our research group has been working for years on the application of methods and concepts of evolution and genetics of molecular populations to the study of pathogenic microorganisms, in what is known as molecular epidemiology. In addition to working on issues of scientific interest, we take problems and return relevant results to the health authorities, achieving an interesting application of a basic biological discipline. In this context, in this project we plan to study a wide prospective collection of isolates of a bacterium of great interest for public health, Klebsiella pneumoniae, to analyse the evolutionary processes that affect its dynamics in the population of the Valencian Community, with special interest in strains resistant to antibiotics. Due to its clinical and public health relevance, we will focus on beta-lactamase producing strains with extended spectrum and/or carbapenemasas. Many of the genes responsible for these phenotypes are located in plasmids that are easily transferred between K. pneumoniae strains and that can also reach it from other species. Therefore, an additional but closely related objective will be the characterization of these plasmids, their mobilization dynamics and, in a more innovative way, their adaptive consequences reflected in the sequences of their genomes. We are particularly interested in the established interactions between plasmid genes and between them and those of the bacterial chromosome, what we know as spystorias. We want to study the relevance of epistasis in the establishment and expansion of multiresistances to antibiotics. This aspect will also be approached with a very different design, of experimental evolution in laboratory cultures. For this we will work with several evolutionary lineages but of epidemiological interest of K. pneumoniae, incorporating an additional level of complexity, the genomic background, in the analysis of epistasis. Our fourth and last objective is based on the results obtained with Treponema pallidum in the current project BFU2014-58656-R and in it we propose to analyse the role of recombination and selection in the epidemic expansion of a T subspecies clone. pallidum pallidum, causal agent of syphilis, which is resistant to azithromycin, an antibiotic that is not commonly used in infections of this bacterium but is frequently used in the treatment of other STIs, such as gonorrhoea and chlamydia. In summary, the project will combine the large-scale study of bacterial genomes, experimental evolution, and molecular and evolutionary epidemiology to study the dynamics of epidemics and mobilization of antibiotic resistance genes in the Valencian Community. The expected results are relevant for regional public health authorities and will contribute to our basic knowledge of the evolution of multi-resistance to antibiotics.
Fernando González Candelas
- Neris García González
- Beatriz Beamud Aranguren
- Marta Pla Díaz
- Carlos Francés Cuesta
- Iván Ansari Toledano
Ministerio de Ciencia, Innovación y Universidades
Fundación para el Fomento de la Investigación Sanitaria y Biomédica de la Comunitat Valenciana (FISABIO)